Research Areas-TEST
Soft Materials
 Soft materials, such as polymers, colloids, surfactants, DNA, and gels are of interest because they spontaneously self-organize into mesoscopic physical structures that can be exploited for nanotechnology or biomedical applications. In the colloids area, she and colleague Orlin Velev are exploring the self-assembly, crystallization and/or gelation of systems of colloid particles with embedded dipoles so as to guide the discovery of advanced materials in the Velev laboratory. Hall has developed a coarse grained model of chitosan, a polysaccharide derived from the shells of crustaceans and is using it to simulate the formation of hydrogels for use in oil spill remediation and drug delivery.
Soft materials, such as polymers, colloids, surfactants, DNA, and gels are of interest because they spontaneously self-organize into mesoscopic physical structures that can be exploited for nanotechnology or biomedical applications. In the colloids area, she and colleague Orlin Velev are exploring the self-assembly, crystallization and/or gelation of systems of colloid particles with embedded dipoles so as to guide the discovery of advanced materials in the Velev laboratory. Hall has developed a coarse grained model of chitosan, a polysaccharide derived from the shells of crustaceans and is using it to simulate the formation of hydrogels for use in oil spill remediation and drug delivery.  With collaborator Tania Betancourt (Texas State University), Hall is using coarse-grained simulations to optimize the design of aptamer-enabled molecularly-responsive hydrogels. A recent focus is the use of simulations to learn the composition of the corona of proteins formed around nanoparticles in the body, a project expected to help us understand nanoparticle toxicity.
With collaborator Tania Betancourt (Texas State University), Hall is using coarse-grained simulations to optimize the design of aptamer-enabled molecularly-responsive hydrogels. A recent focus is the use of simulations to learn the composition of the corona of proteins formed around nanoparticles in the body, a project expected to help us understand nanoparticle toxicity.
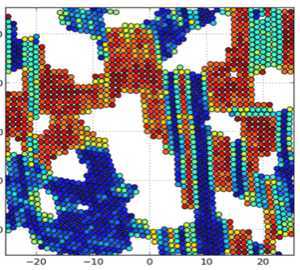
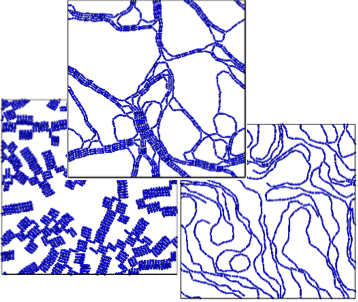 Self-assembly of Dipolar Colloids: David Rutkowski and Ryan Maloney
Self-assembly of Dipolar Colloids: David Rutkowski and Ryan Maloney
Discontinuous molecular dynamics and Monte Carlo simulations of 2-d systems of dipolar rods and of spheres with dipoles offset from the center line are conducted. Phase diagrams in the temperature – area fraction plane outlining the boundaries between fluid, string-fluid and percolated states are determined. The emphasis here is on network formation.
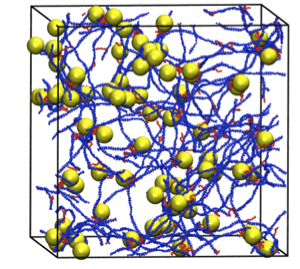 Simulations of Chitosan Assembly: Steven Benner
Simulations of Chitosan Assembly: Steven Benner
Chitosan is a versatile biopolymer with potential applications ranging from drug delivery and wound healing to flocculation of oil/water emulsions. Coarse-grained models of chitosan and hydrophobically modified chitosan have been developed that allow simulation via discontinuous molecular dynamics of the assembly of chitosan to form networks that encapsulate drug molecules.
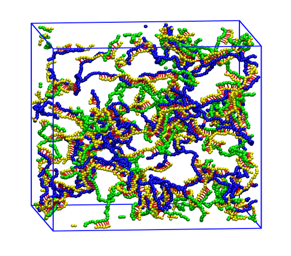 Aptamer-crosslinked Hydrogels: Kye Won Wang
Aptamer-crosslinked Hydrogels: Kye Won Wang
Aptamer-crosslinked hydrogels are being developed in the laboratory of our collaborator, T. Betancourt, for use as stimuli-responsive drug delivery systems. Coarse grained molecular dynamics simulations are being used to analyze their formation, drug loading capacity and degradation.
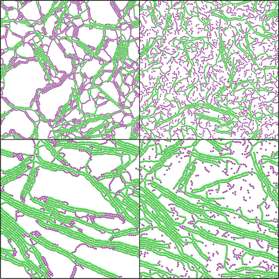 Self-assembling Mixtures of Dipolar Rods and Spheres: Ryan Maloney
Self-assembling Mixtures of Dipolar Rods and Spheres: Ryan Maloney
Discontinuous molecular dynamics simulations of mixtures of dipolar rods and spheres reveal that they have much more complex behavior than is seen for either single component systems. Brownian dynamics simulations of mixtures of active and dipolar discs are also being conducted. These mixtures offer a potential path to creating new, temperature responsive, bifunctional materials
 Underwater Adhesives: Amelia Chen and Qing Shao
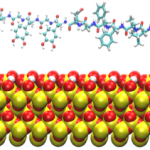
Underwater Adhesives: Amelia Chen and Qing Shao
Studies on underwater organisms such as mussels and barnacles have shown that their natural glues involve two key components: L-3,4,-dihydroxyphenylalanine (L-DOPA) and amyloid nanostructures. Simulations are being used to investigate the behavior of L-DOPA and amyloid-forming peptides near surfaces, the goal being to design of synthetic, generic underwater adhesives.
 Protein Nanoparticle Interactions: Qing Shao
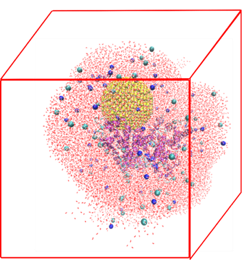
Protein Nanoparticle Interactions: Qing Shao
We investigated the allosteric effects of nanoparticles on proteins using a multiscale modeling approach and found that nanoparticles can change the properties of proteins even on regions distant from their binding site. This finding opens new ways to design nanoparticles for medical applications.
Protein Aggregation
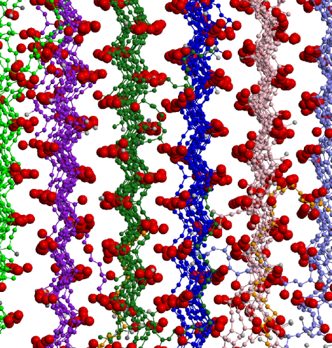 Protein aggregation is a cause or associated symptom of a number of neurodegenerativediseases, including Alzheimer’s, Parkinson’s and the prion diseases. Hall and her students are using computer simulation to investigate the formation of ordered protein aggregates, called fibrils, which are invariably found in the brains of victims of these diseases. A coarse-grained protein model, PRIME20, has been developed that enables the simulation of specific amyloidogenic peptides when used with discontinuous molecular dynamics (DMD), a fast alternative to traditional molecular dynamics. Hall was among the first to tackle the protein aggregation of a large system of proteins starting from a random configuration of monomers and culminating in fibrillar structures. As part of collaboration with experimental biophysicist, Sheena Radford, and chemist, Andrew Wilson, at Leeds University, she is simulating the early oligomerization and fibrillation of beta amyloid, the Alzheimer’s peptide, and evaluating the impact of selected aggregation inhibitors. Finally, she and collaborators Anant Paravastu (Georgia Tech) and Gregory Hudalla (University of Florida) are investigating the co-assembly of peptides used as recombinant protein fusion tags for integrating enzymes into supramolecular hydrogels.
Protein aggregation is a cause or associated symptom of a number of neurodegenerativediseases, including Alzheimer’s, Parkinson’s and the prion diseases. Hall and her students are using computer simulation to investigate the formation of ordered protein aggregates, called fibrils, which are invariably found in the brains of victims of these diseases. A coarse-grained protein model, PRIME20, has been developed that enables the simulation of specific amyloidogenic peptides when used with discontinuous molecular dynamics (DMD), a fast alternative to traditional molecular dynamics. Hall was among the first to tackle the protein aggregation of a large system of proteins starting from a random configuration of monomers and culminating in fibrillar structures. As part of collaboration with experimental biophysicist, Sheena Radford, and chemist, Andrew Wilson, at Leeds University, she is simulating the early oligomerization and fibrillation of beta amyloid, the Alzheimer’s peptide, and evaluating the impact of selected aggregation inhibitors. Finally, she and collaborators Anant Paravastu (Georgia Tech) and Gregory Hudalla (University of Florida) are investigating the co-assembly of peptides used as recombinant protein fusion tags for integrating enzymes into supramolecular hydrogels.
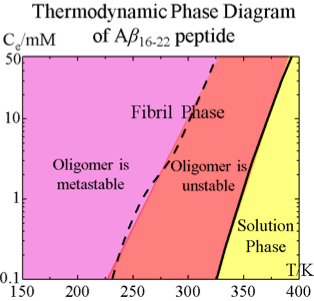
Aggregation of Aβ16-22 Peptides : Yiming Wang
Aβ16-22 is a central hydrophobic fragment of Aβ protein. It’s aggregation kinetics is investigated by applying discontinuous molecular dynamics with the PRIME20 protein model. The thermodynamics of Aβ16-22 is revealed by constructing an equilibrium phase diagram in the temperature – concentration plane.

Inhibition of Aβ17-36 Aggregation : Yiming Wang
Some naturally occurring polyphenols (vanillin, resveratrol, curcumin and EGCG) can inhibit Aβ fibril formation and reduce neuron cell toxicity in vitro. Coarse-grained simulations were used to investigate their inhibition mechanism, helping us to understand the structure-function relationship underlying well-known amyloid inhibitors and design new ones.

Co-assembly of CATCH Peptides: Qing Shao
Peptide co-assembly is emerging as a new process to prepare nanomaterials. We are collaborating with experimentalists Gregory Hudalla (U Florida) and Anant Paravastu ( Georgia Tech) to understand the fundamentals that control peptide co-assembly and design high-performance co-assembling peptide systems.
Protein Design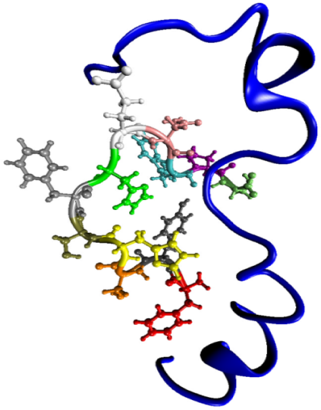
Short peptides that bind to specific biomolecules have many uses, including as biosensors, diagnostics, therapeutics, and affinity ligands for bioseparations. The Hall group has been developing a computational search algorithm to design peptide sequences that can bind to specific RNA- or protein-based targets. The method has been applied to design a 15-amino-acid peptide to recognize human lysine tRNA species (tRNALys3), a primer for HIV reverse transcription, as part of a collaboration with Paul Agris at Albany University. In a project with researchers at the Air Force Research Laboratory, peptides are being designed to produce high-sensitivity detectors of stress-related biomarkers like cardiac troponin I (cTnl), neuropeptide Y, and cortisol.
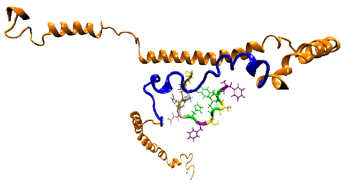 Designing Peptides to Recognize Heart Attack Biomarkers: Xingqing Xiao
Designing Peptides to Recognize Heart Attack Biomarkers: Xingqing Xiao
A search algorithm was developed to design peptides that bind to specific biomolecular targets including cardiac troponin I, a heart attack biomarker in collaboration with R. Naik and P. Mirau at the Air Force Research lab. One in-silico peptide has ~16× higher affinity than the phage-display discovered peptide and a much lower limit of detection. The algorithm is being used to design sensors for cortisol.

Peptide Search Algorithms to Block Recruitment of tRNA-Lys3 by HIV: Xingqing Xiao
Blocking the recruitment of human tRNALys3, a primer for reverse transcription, by HIV has the potential to interfere with the virus life cycle. Our peptide search algorithm was used to design 15-residue peptides which could bind selectively to the anticodon stem and loop on tRNALys3 . When synthesized and characterized experimentally in the lab of our collaborator Paul Agris (Albany University) one of our designed peptides bound to the tRNALys3 with 10-fold higher affinity and specificity than the best phage-display identified peptide.